Current Projects
Accelerating peptide drug discovery & design.The peptide space is vast, but the regions occupied by biologically active peptides account for a limited set of sequences folding into privileged structures. The advent of artificial intelligence (AI) in life sciences has led to the creation of machine learning (ML) models capable of predicting sequence-structure-function relationships (that may not be readily apparent) and generating new proteins with the desired characteristics. Predictive models infer the relationships between the protein sequences and their biological functions (e.g., thermostability, microbial inhibition, protein binding affinity). Generative models learn meaningful representations from sequences or structures to create proteins that resemble their native counterparts. Successful applications of ML-guided protein design have been documented worldwide for protein binders, antigen-specific monoclonal antibodies, protein families, and enzymes. My group devises predictive models and bioinformatic pipelines to effectively screen for peptide sequences from sizable public repositories or proteo-transcriptomic datasets. Our efforts focused on antimicrobial peptides and venom toxins. Both groups of bioactive peptides are well-documented in curated databases, private repositories, and in-house datasets. Our current research efforts focus on (i) discovering novel peptides from untapped extant species using prior knowledge and (ii) engineering novel peptides with desired characteristics in multi-objective optimisation.

Designing peptide therapeutics with explainable AI.
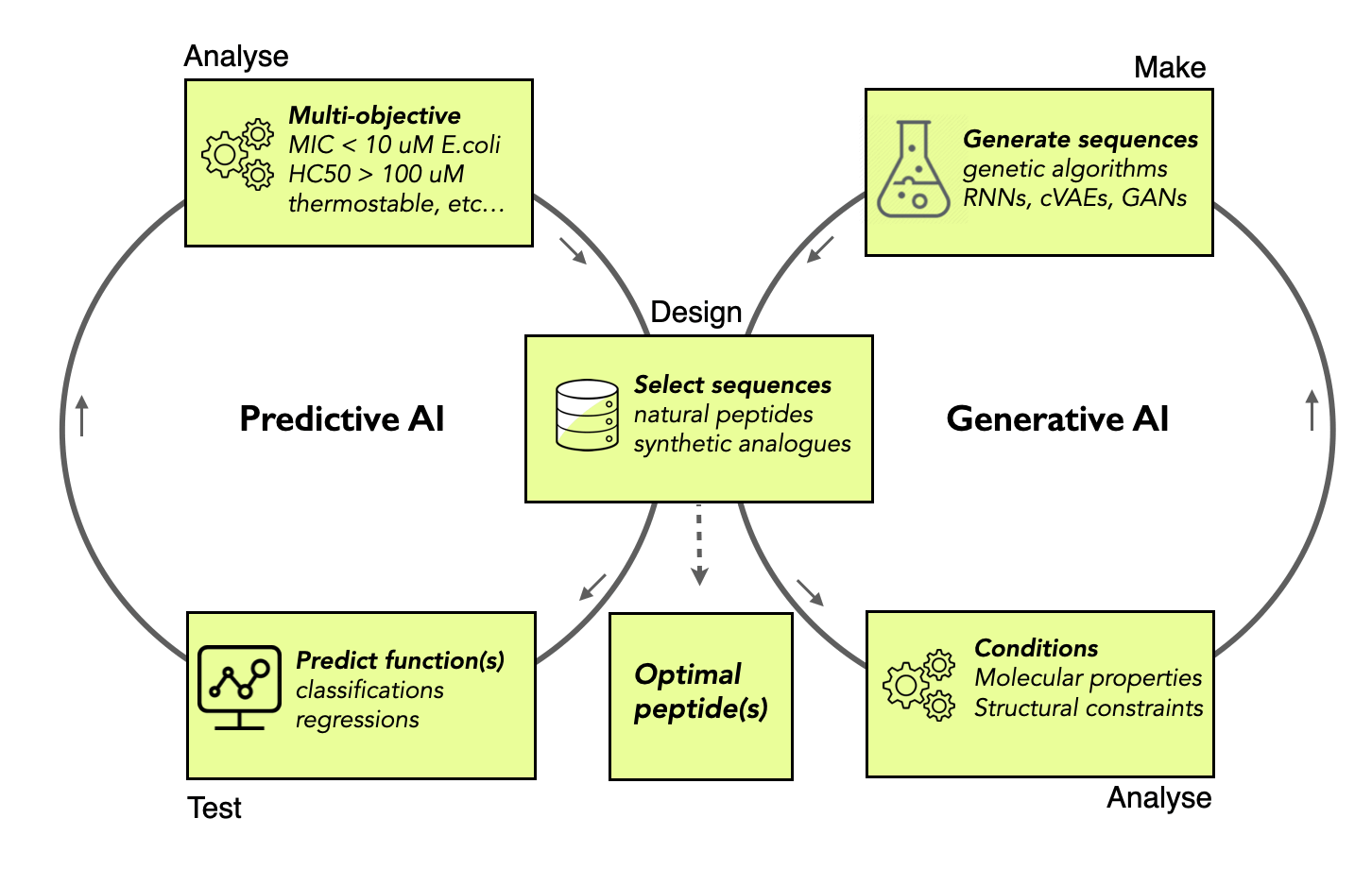
An essential step in peptide drug design is finding the trade-off between the extraordinary biological activities and highly desirable pharmacokinetic properties of peptide candidates while reducing their undesirable side effects (e.g., lack of metabolic stability, lability during storage, poor oral bioavailability, and unwanted toxicity) – a multi-objective optimization. Conventional approaches operate iteratively through a series of modifications, followed by their physical or biological evaluations – a Design-Make-Test-Analyse (DMTA) cycle. The peptide space is vast - investigating all functionally interesting analogues is not practical and is an expensive task. Implementing a computational DMTA cycle speeds up the design process by narrowing the list of peptide candidates for synthesis and biological evaluations. To minimise the impact that structural factors might have upon biological prediction or sequence generation, researchers voluntarily select sequences based on structural or evolutionary constraints, which presumably adopt the same tridimensional structure(s). Our recent efforts in discovering novel bioactive peptides also revealed that computational designers have so far capitalised on ML architectures to predict and generate novel α-helical non-hemolytic AMPs. These findings indicate that current ML models do not generalize well and are unfair towards minor structural classes, limiting the prospect of intellectual property.
Producing our AI-designed peptides.
Accessing peptide libraries is crucial to understanding and validating our AI-driven hypotheses about complex sequence-structure-function relationships. We have been exploring ways to produce our peptides through solid-phase peptide synthesis (SPPS) and protein expression in biological systems. During my postdoctoral stays at the University of Queensland, I synthesised peptide analogues using both manual and automated SPPS. Our current efforts have shifted towards the production of peptides in biological expression systems; bacteria (Escherichia coli), yeast (Pichia pastoris), and tobacco plant (Nicotiana benthamiana) due to the institutional facilities in place. Both strategies are ideal for producing a handful of bioactive peptides. However, SPPS and heterologous expression in biological systems have their own limitations when it comes to generating large numbers of peptides and proteins.
Past Projects
Developed Reac2Vac and DeepPlay, two application programming interfaces.Insight Data, Boston USA [2016-2017] [Reac2Vac Github] [DeepPlay Github]
Designed peptide inhibiting protein-protein interactions in use for acute myeloid leukemia.UQ Pharmacy Australia Centre of Excellence and Protagonist Therapeutics, Brisbane Australia [2015].
Designed novel potent GLP-1R peptide agonists in application to type 2 diabetes.UQ Institute for Molecular Bioscience, Brisbane Australia and Pfizer, Cambridge and Groton USA [2012-2015].
Studied the physicochemical properties of bioavailable cyclic peptides and macrocycles.UQ Institute for Molecular Bioscience, Brisbane Australia and Pfizer, Cambridge and Groton USA [2012-2015].
High-throughput screening natural products for antimalarial agents.UQ Institute for Molecular Bioscience and Queensland Institute for Medical Research, Brisbane Australia [2012].
Developed ML models to predict the blood-brain barrier permeability applied to natural products.UQ Institute for Molecular Bioscience, Brisbane Australia and Noscira, Madrid Spain [2008-2012].
Repositioned in-house natural product library to discover novel kinase inhibitors.UQ Institute for Molecular Bioscience, Brisbane Australia and Noscira, Madrid Spain [2008-2012].
Discovered novel kinase inhibitors from marine natural product chemistry for Alzheimer's disease.UQ Institute for Molecular Bioscience, Brisbane Australia and Noscira, Madrid Spain [2008-2012].
Hit/Lead optimization of natural product-derived proteasome inhibitors for cancer treatments.
Institut de Sciences and Technologies du Médicament de Toulouse (ISTMT),
Centre National de la Recherche Scientifique - Pierre Fabre Group, Toulouse France [2007-2008].
Department of Chemistry, University of Aberdeen, Aberdeen Scotland [2006].
Designed small molecules with liquid crystal propertiesDepartment of Chemistry, University of Hull, Hull England [2005].